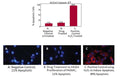
- Prepare samples and controls
- Reconstitute Magic Red with 100 or 400 µL DMSO
- Dilute Magic Red 1:5 with diH2O
- Add diluted Magic Red at 1:15-1:30 to suspension cells
- Incubate while protected from light.
- Watch color start to develop within 15 minutes of addition of Magic Red
- If desired, label with additional stains, such as Hoechst, DAPI, or an antibody.
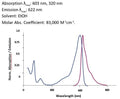
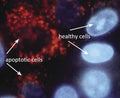
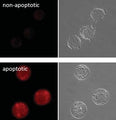
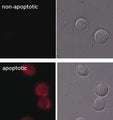
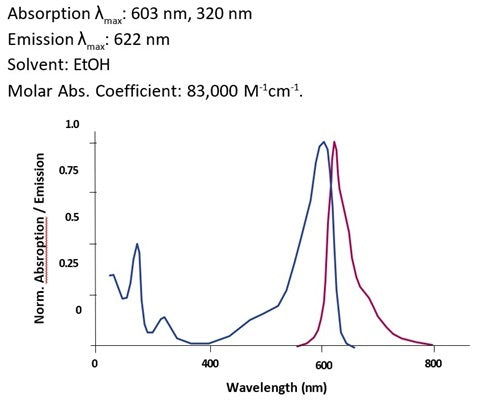
- Analyze with a fluorescent microscope, fluorescent plate reader, or flow cytometer. Magic Red has a maximum excitation at 592 nm and emission at 628 nm. Good fluorescence images can be obtained at 510-560 nm excitation and >610 nm emission.
If working with adherent cells, please see the manual for additional protocols.
Product Specific References
| PMID | Publication |
| 39487138 | Sun, F, et al. 2024. Increased DNA damage in full-grown oocytes is correlated with diminished autophagy activation. Nature Communications, 9463. |
| 36632900 | Benavides, R.A.S., et al. 2023. The HL-60 human promyelocytic cell line constitutes an effective in vitro model for evaluating toxicity, oxidative stress and necrosis/apoptosis after exposure to black carbon particles and 2.45 GHz radio frequency. The Science of the total environment, 161475. |
| 37036227 | Tong, J., et al. 2023. Measuring Caspase Activity Using a Fluorometric Assay or Flow Cytometry. Journal of visualized experiments : JoVE. |
| 37257963 | Procházková, M., et al. 2023. Novel Förster Resonance Energy Transfer probe with quantum dot for a long-time imaging of active caspases inside individual cells. Analytica chimica acta, 341334. |
| 36346836 | Ndinyanka Fabrice, T., et al. 2022. An evolutionarily conserved coronin-dependent pathway defines cell population size. Science signaling, eabo5363. |
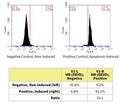
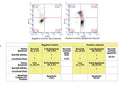
Question: The customer wish to combine in the same sample the Magic Red staining and Annexin V staining. The analysis of the samples will be performed on a flow cytometer. In the manual, page 10, section: single satin Flowcytometry, after the MR incubation there is no wash step. Since the Annexin V staining procedure can be performed only with a staining buffercontaining calcium, can the MR be performed in this buffer instead of the cells media? So that the customer will begin with the MR staining and in the last 15 min she will add the Annexin V. Another option can be perhaps to stain with the MR, wash and replace it to the Annexin V buffer for future 15 min staining. Will the MR “survive” a centrifuge?
Answer: We have not evaluated our Magic Red substrates in combination with Annexin V, so unfortunately can’t say with certainty that this will work. However, that said, we would recommend against the proposed sequence where the customer stains with MR, washes, resuspends in Annexin V buffer, and then completes the Annexin V staining step. MR substrate is not covalently coupled in the cell and the signal would be washed away in the centrifugation step. The customer can try completing Annexin V staining and then carrying our MR staining afterwards in the Annexin V buffer. We would once again caution that the use of this technique will need to be evaluated experimentally. We recommend including positive and negative single stain controls for interpretation of the results.