- Prepare samples and controls.
- Dilute 10X Cellular Wash Buffer 1:10 with diH20.
- Reconstitute FLICA with 50 µL DMSO.
- Dilute FLICA 1:5 by adding 200 µL PBS.
- Add diluted FLICA to each sample at 1:30-1:60 (e.g. spike at 1:30 by adding 10 µL to 290 µL cultured cells).
- Incubate approximately 1 hour.
- Remove media and wash cells 3 times: add 1X Cellular Wash Buffer and spin cells.
- If desired, label with additional stains, such as Hoechst, DAPI, or an antibody.
- If desired, fix cells.
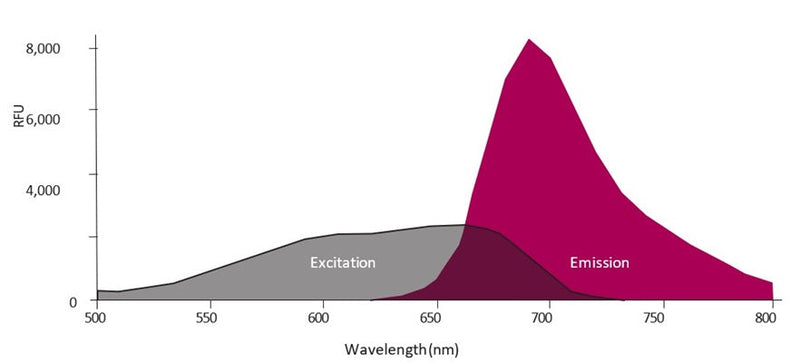
- Analyze with a fluorescence microscope or flow cytometer. FLICA 660 excites at 660 nm and emits at 680-690 nm.
Product Specific References
| PMID | Publication |
| 39908772 | Wu, Z, et al. 2025. Qufeng epimedium decoction alleviates rheumatoid arthritis through CYLD-antagonized NF-kB activation by deubiquitinating Sirt1. Immunobiology, 152875. |
| 39878051 | Mao, S, et al. 2025. Design, Synthesis and Anti-Inflammatory Evaluation of 3-Substituted 5-Amidobenzoate Derivatives as Novel P2Y14 Receptor Antagonists via Structure-Guided Molecular Hybridization. Journal of Medicinal Chemistry, . |
| 39706368 | Martins-Gonçalves, R, et al. 2024. Platelet-neutrophil aggregate formation induces NLRP3 inflammasome activation in vaccine-induced thrombotic thrombocytopenia. Journal of Thrombosis and Haemostasis, . |
| 39738348 | Chen, R, et al. 2024. Total glucosides of paeony ameliorates chemotherapy-induced neuropathic pain by suppressing microglia pyroptosis through the inhibition of KAT2A-mediated p38 pathway activation and succinylation. Scientific Reports, 31875. |
| 39568027 | Zhang, Y., et al. 2024. Neutrophil extracellular traps facilitate liver inflammation/fibrosis progression by entering macrophages and triggering AIM2 inflammasome-dependent pyroptosis. Cell Communication and Signaling, 556. |
| 39563448 | Cheng, J., et al. 2024. NEK7 induces lactylation in Alzheimer's disease to promote pyroptosis in BV-2 cells. Molecular Brain, 81. |
| 39550359 | Anderson, M.J., et al. 2024. Broad-spectrum inflammasome inhibition by thiomuscimol. Cell Death & Disease, 470. |
| 39532893 | He, Y., et al. 2024. Mechanics-activated fibroblasts promote pulmonary group 2 innate lymphoid cell plasticity propelling silicosis progression. Nature Communications, 9770. |
| 39486414 | Hu, Y, et al. 2024. Identification of two repurposed drugs targeting GSDMD oligomerization interface I to block pyroptosis. Cell Chemical Biology, . |
| 39455437 | Kong, R, et al. 2024. The role of Gαq in regulating NLRP3 inflammasome activation. Inflammation Research, . |
| 39377494 | Shang, C, et al. 2024. Functional evaluation of TMEM176B and its predictive role for severe respiratory viral infection through integrated analysis of single-cell and bulk RNA-sequencing. Journal of Medical Virology, e29954. |
| 39312228 | Wackerbarth, L.M., et al. 2024. Neonatal neutrophils exhibit reduced NLRP3 inflammasome activation. Journal of Leukocyte Biology, . |
| 39298552 | Ogasawara-Nosoko, M., et al. 2024. COMMD5 counteracts cisplatin-induced nephrotoxicity by maintaining tubular epithelial integrity and autophagy flux. American Journal of Physiology. Renal Physiology, . |
| 39240187 | Hsieh, L.L., et al. 2024. Bystander monocytic cells drive infection-independent NLRP3 inflammasome response to SARS-CoV-2. mBio, e0081024. |
| 39232171 | Narasimhan, H., et al. 2024. An aberrant immune-epithelial progenitor niche drives viral lung sequelae. Nature, . |
| 39112714 | Cao, J., et al. 2024. DNA-sensing inflammasomes cause recurrent atherosclerotic stroke. Nature, . |
| 39097915 | Zhang, H., et al. 2024. Astragaloside IV promotes the pyroptosis of airway smooth muscle cells in childhood asthma by suppressing HMGB1/RAGE axis to inactivate NF-κb pathway. Autoimmunity, 2387100. |
| 39044033 | Song, Z, et al. 2024. An NAD+-dependent metabolic checkpoint regulates hematopoietic stem cell activation and aging. Nature aging, . |
| 39040105 | Akiyama, M, et al. 2024. An early regulatory mechanism of hyperinflammation by restricting monocyte contribution. Frontiers in immunology, 1398153. |
| 39033542 | Ma, H, et al. 2024. Gasdermin D Mediated Mitochondrial Metabolism Orchestrate Neurogenesis Through LDHA During Embryonic Development. Advanced science (Weinheim, Baden-Wurttemberg, Germany), e2402285. |
| 39032599 | Cao, Y, et al. 2024. Caspase-1 Inhibition Ameliorates Photoreceptor Damage Following Retinal Detachment by Inhibiting Microglial Pyroptosis. The American journal of pathology, . |
| 39002871 | Wang, M, et al. 2024. Apabetalone, a BET protein inhibitor, inhibits kidney damage in diabetes by preventing pyroptosis via modulating the P300/H3K27ac/PLK1 axis. Pharmacological research, 107306. |
| 38996778 | Shen, L, et al. 2024. Macrophage α7nAChR alleviates the inflammation of neonatal necrotizing enterocolitis through mTOR/NLRP3/IL-1β pathway. International immunopharmacology, 112590. |
| 38897422 | Russell-Guzmán, J, et al. 2024. Activation of the ROS/TXNIP/NLRP3 pathway disrupts insulin-dependent glucose uptake in skeletal muscle of insulin-resistant obese mice. Free radical biology & medicine, 187-198. |
| 38874515 | Liu, W, et al. 2024. Discovery of N-Substituted Acetamide Derivatives as Promising P2Y14R Antagonists Using Molecular Hybridization Based on Crystallographic Overlay. Journal of medicinal chemistry, 10233-10247. |
| 38510374 | Wang, X, et al. 2024. Lactobacillus Plantarum Promotes Wound Healing by Inhibiting the NLRP3 Inflammasome and Pyroptosis Activation in Diabetic Foot Wounds. Journal of inflammation research, 1707-1720. |
| 38530976 | Su, X.J., et al. 2024. Exosomes Derived from Caerulein-Stimulated Pancreatic Acinar Cells Mediate Peritoneal Macrophage M1 Polarization and Pyroptosis via an miR-24-3p/MARCH3/NLRP3 Axis in Acute Pancreatitis. Pancreas. |
| 38428396 | Wang, Q., et al. 2024. The CARD8 inflammasome dictates HIV/SIV pathogenesis and disease progression. Cell, 1223-1237.e16. |
| 38341879 | Zhang, B., et al. 2024. 1,2-Dichloroethane induces testicular pyroptosis by activating piR-mmu-1019957/IRF7 pathway and the protective effects of melatonin. Environment international, 108480. |
| 38325020 | Wu, X., et al. 2024. Xuebijing improves inflammation and pyroptosis of acute lung injury by up-regulating miR-181d-5p-mediated SPP1 inactivation. Clinics (Sao Paulo, Brazil), 100336. |
| 38189627 | Pu, C., et al. 2024. Low-Dose Chemotherapy Preferentially Shapes the Ileal Microbiome and Augments the Response to Immune Checkpoint Blockade by Activating AIM2 Inflammasome in Ileal Epithelial Cells. Advanced science (Weinheim, Baden-Wurttemberg, Germany), e2304781. |
| 37998361 | Spurlock, M., et al. 2023. The Inflammasome-Dependent Dysfunction and Death of Retinal Ganglion Cells after Repetitive Intraocular Pressure Spikes. Cells. |
| 37894824 | Leinardi, R., et al. 2023. Distinct Pro-Inflammatory Mechanisms Elicited by Short and Long Amosite Asbestos Fibers in Macrophages. International journal of molecular sciences, . |
| 37749820 | Wang, Q, et al. 2024. Tauroursodeoxycholic acid protects Schwann cells from high glucose-induced cytotoxicity by targeting NLRP3 to regulate cell migration and pyroptosis. Biotechnology and Applied Biochemistry, 28-37. |
| 37123216 | Li, H., et al. 2023. Heme oxygenase‑1 inhibits renal tubular epithelial cell pyroptosis by regulating mitochondrial function through PINK1. Experimental and Therapeutic Medicine, 213. |
| 37093343 | Liang, M.Q., et al. 2023. LncRNA SNHG3 Promotes Sevoflurane-Induced Neuronal Injury by Activating NLRP3 via NEK7. Neurochemical research. |
| 37182453 | Yan, C., et al. 2023. Endoplasmic reticulum stress promotes caspase-1-dependent acinar cell pyroptosis through the PERK pathway to aggravate acute pancreatitis. International immunopharmacology, 110293. |
| 37356296 | Mikolajczyk-Martinez, A., et al. 2023. Unraveling the Role of Type 1 Fimbriae in Salmonella Pathogenesis: Insights from a Comparative Analysis of Salmonella Enteritidis and Salmonella Gallinarum. Poultry Science, 102833. |
| 37602503 | Zou, H., et al. 2023. C/EBPβ isoform-specific regulation of podocyte pyroptosis in lupus nephritis-induced renal injury. The Journal of pathology. |
| 37556724 | Torraca, V., et al. 2023. Shigella serotypes associated with carriage in humans establish persistent infection in zebrafish. The Journal of infectious diseases. |
| 35151118 | Zhang, C., et al. 2022. Targeting NLRP3 Signaling by a Novel-designed Sulfonylurea Compound for Inhibition of Microglial Inflammation. Bioorganic & Medicinal Chemistry, 116645. |
| 35180562 | Zeng, M., et al. 2022. LncRNA ROR promotes NLRP3-mediated cardiomyocyte pyroptosis by upregulating FOXP1 via interactions with PTBP1. Cytokine, 155812. |
| 35721108 | Wu, X., et al. 2022. Emodin Ameliorates Acute Pancreatitis-Associated Lung Injury Through Inhibiting the Alveolar Macrophages Pyroptosis. Frontiers in pharmacology, 873053. |
| 35835255 | Bertoni, A., et al. 2022. Spontaneous NLRP3 inflammasome-driven IL1-β secretion is induced in severe COVID-19 patients and responds to anakinra treatment. The Journal of allergy and clinical immunology. |
| 35761192 | Lan, J., et al. 2022. WTAP-mediated N6-methyladenosine modification of NLRP3 mRNA in kidney injury of diabetic nephropathy. Cellular & molecular biology letters, 51. |
| 36129987 | Duhalde Vega, M., et al. 2022. PD-1/PD-L1 blockade abrogates a dysfunctional innate-adaptive immune axis in critical β-coronavirus disease. Science advances, eabn6545. |
| 36155068 | Shu, L., et al. 2022. PHLDA1 promotes sevoflurane-induced pyroptosis of neuronal cells in developing rats through TRAF6-mediated activation of Rac1. Neurotoxicology, 140-151. |
| 36357533 | Clark, K.M., et al. 2022. Chemical inhibition of DPP9 sensitizes the CARD8 inflammasome in HIV-1-infected cells. Nature chemical biology. |
| 36429008 | Liu, Y., et al. 2022. S100A8-Mediated NLRP3 Inflammasome-Dependent Pyroptosis in Macrophages Facilitates Liver Fibrosis Progression. Cells, 3579. |
| 33667846 | Lu, R., et al. 2021. Design, synthesis and evaluation of 3-amide-5-aryl benzoic acid derivatives as novel P2Y14R antagonists with potential high efficiency against acute gouty arthritis. European Journal of Medicinal Chemistry, 113313. |
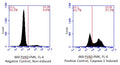
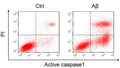
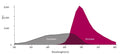
Question: FLICA 660 optimally excites at 660 nm and has a peak emission at 685-690 nm, and Propidium Iodide excites at 615nm. Can I use the FLICA 660 and the Propidium Iodide to detect pyroptosis by Flow cytometry?
Answer: We have done some two-color panels pairing FLICA 660 with green emission fluors, however, we have not evaluated it alongside Propidium Iodide. That said, despite emission spectra overlap between Propidium Iodide and 660-YVAD-FMK I believe it should be possible to resolve the red vs far red fluors with appropriate compensation. I wanted to make you aware of Green Live/Dead Stain, ICT’s membrane impermeant green fluorescent vital stain for differentiating live and dead cells. This product performs similarly to Propidum Iodide, however, due to its green emission properties, it is compatible with our FLICA 660 products without the need for compensation.